About nanoPASS
Goal
nanoPASS will develop and validate a technology for resource efficient animal free prediction of nanomaterial and advanced material induced long-term adverse outcomes and demonstrate its applicability in regulatory and industrial setting.
Objectives
nanoPASS has four main objectives:
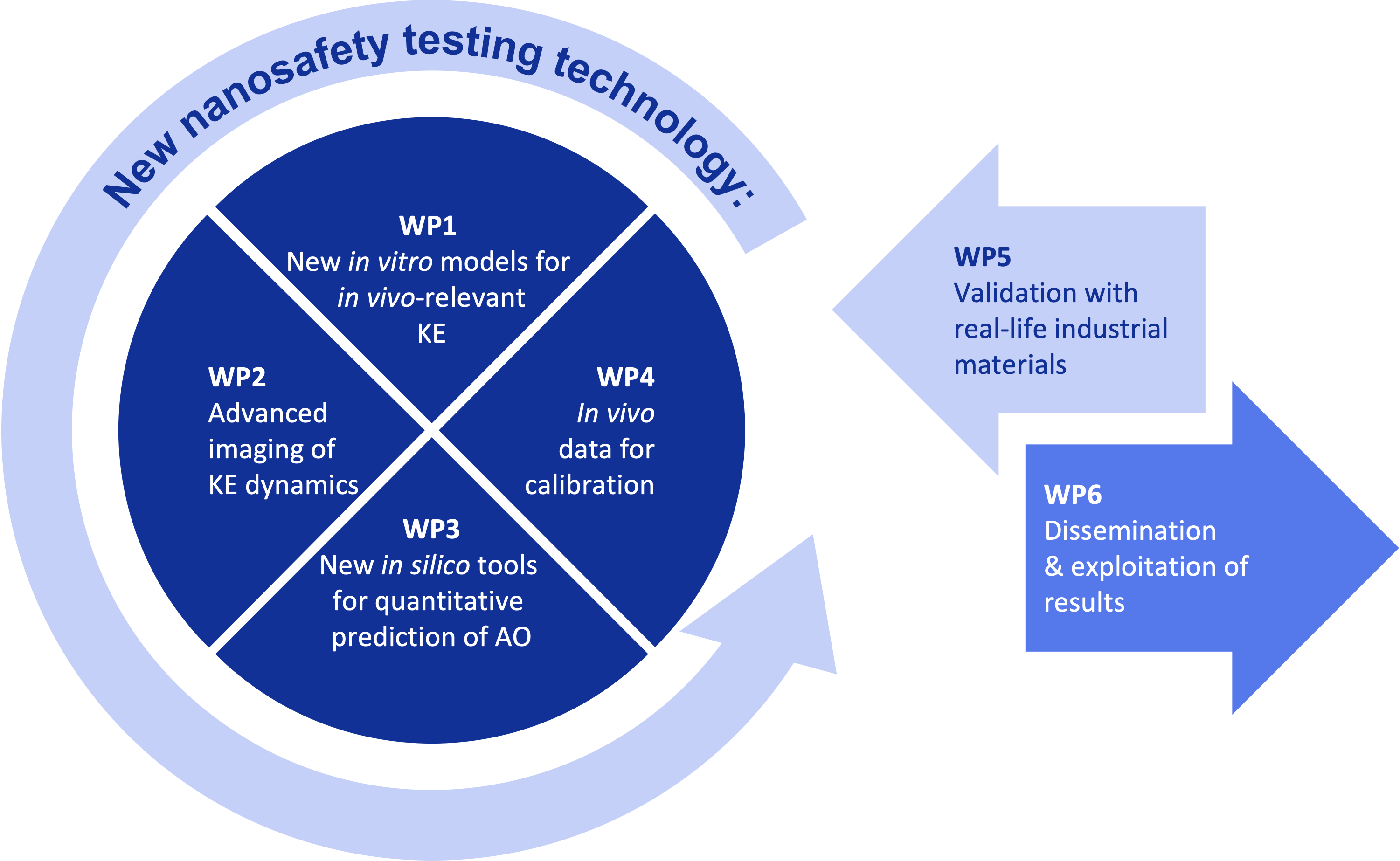
Objective 1: Develop new in vitro models relevant for prediction of in vivo adverse outcomes (AOs). We will work on in vitro systems that recapitulate in vivo-relevant key events (KEs) for the chosen AOs, which enable fast and high-throughput testing.
Objective 2: Identify suitable sets of methods to track the dynamics of key events, particularly with high-resolution timelapse microscopy techniques and quantitative analysis.
Objective 3: Develop in silico models for quantitative prediction of in vivo adverse outcomes from in vitro data, incorporating automated connection of new KEs into quantitative AOPs (qAOPs).
Objective 4: Calibrate and validate the in vitro/in silico adverse outcome prediction models with in vivo data for 40+ benchmark materials and diverse industrial materials (cement, nanoplastics, electronics waste material, and advanced materials for catalysis and medical applications), collected at different stages of their life cycle.
The objectives will be demonstrated for the 4 chosen adverse outcomes: reduced lung function (AOP302), chronic inflammation (KE in AOP173, fibrosis), cardiovascular disease (AOP237), and neurodegeneration.
Impacts
Science: new early KE and their relationships in AOP-Wiki, and FAIR data in eNanoMapper, will improve understanding of NMs’ mechanisms-of-action.
Industry: new tools for fast safety testing, desired by material developers and producers in multibillion markets: construction, chemicals, electronics, medical implants, etc.; new market opportunities for animal-free testing.
Regulators: our application for an OECD test guideline will pave the way towards affordable high-throughput tests that can be widely recommended or mandated.
Society: our industrial demo of hazard assessment will highlight how to better protect consumers, workforce, and environment.
Methodology
We will search for bridging KEs both top-down from in vitro data and bottom-up from in vivo data, using complementary methods, particularly live-cell & intravital microscopy and omics.